The Epigenetic Revolution: A Comprehensive Report on the Science, Clinical Landscape, and Future of Programmable Gene Regulation
Executive Summary
The therapeutic manipulation of the human genome has entered a transformative era. For decades, the conceptual framework of genetic medicine was defined by the binary nature of the DNA sequence: genes were either present or absent, mutated or wild-type. The advent of programmable nucleases, culminating in the Nobel Prize-winning CRISPR-Cas9 technology, provided the tools to edit this "hardware" of life. However, a parallel and perhaps more nuanced revolution is rapidly maturing: the ability to edit the "software" of the cell—the epigenome.
Epigenetic editing, the precise, programmable modulation of gene expression without altering the underlying DNA sequence, represents a paradigm shift from irreversible genetic surgery to tunable, reversible transcriptional control. By leveraging the cell's endogenous regulatory mechanisms—methylation, histone modification, and chromatin remodeling—this technology offers solutions to the most persistent challenges of traditional gene editing: the risks of genotoxicity, the difficulty of multiplexing, and the inability to address conditions requiring subtle regulation rather than binary ablation.
This report provides an exhaustive analysis of the current state of epigenetic editing. It deconstructs the biological mechanisms that underpin epigenetic memory, contrasts the modality with traditional gene editing, evaluates the complex risk profiles including transgenerational inheritance, and details the rapidly evolving clinical landscape led by pioneers such as Tune Therapeutics, Epic Bio, and nChroma Bio. Furthermore, it projects the future applications of this technology, particularly in the realms of polygenic disease management, agricultural biotechnology, and the ambitious pursuit of reversing cellular aging.
Chapter 1: The Epigenetic Landscape
1.1 The Biological Substrate of Gene Regulation
To fully appreciate the mechanism and potential of epigenetic editing, one must first understand the complex biological substrate upon which it acts. The human genome is often compared to a library of instructions, but this analogy fails to capture the dynamic nature of DNA regulation. A more accurate comparison is an operating system where access to specific files (genes) is strictly controlled by a sophisticated system of permissions and access rights. This system is the epigenome.
The epigenome consists of chemical modifications to the DNA and its associated proteins that dictate the accessibility of genetic information. While every cell in the human body contains the exact same DNA sequence (the genotype), epigenetic marks determine whether a cell functions as a neuron, a hepatocyte, or a cardiomyocyte (the phenotype).1 This regulation is dynamic, reversible, and responsive to environmental cues, yet stable enough to be maintained through cellular division—a property known as cellular memory.2
1.2 The Trinity of Epigenetic Mechanisms
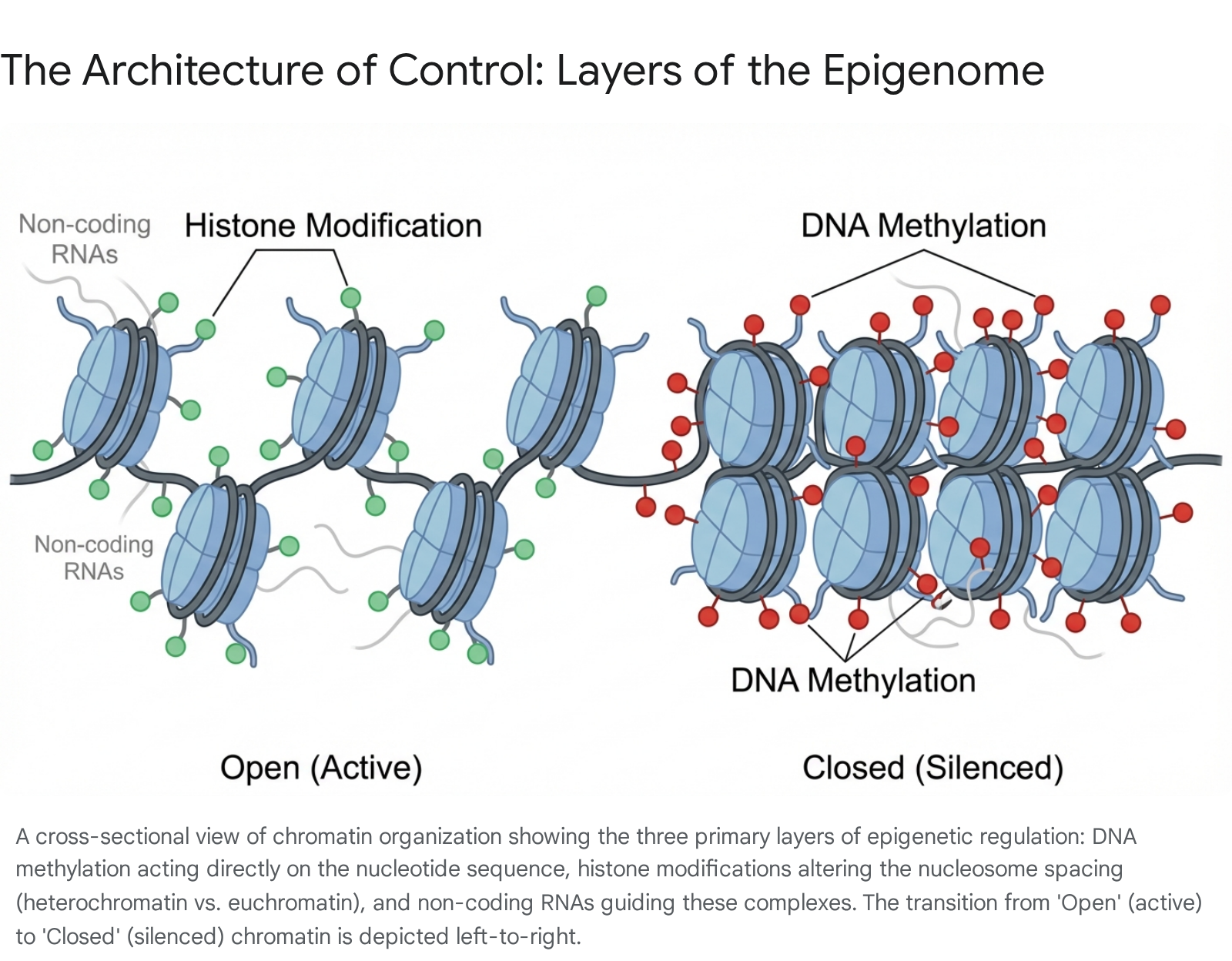
Epigenetic regulation is achieved through three primary, interconnected mechanisms that work in concert to control the chromatin state.
1.2.1 DNA Methylation: The Silencing Lock
The most stable and well-characterized epigenetic mark is DNA methylation. Biochemically, this involves the covalent addition of a methyl group (-CH3) to the 5-carbon position of the cytosine ring, forming 5-methylcytosine (5mC).1 In mammals, this modification predominantly occurs at cytosines that are immediately followed by guanines, known as CpG dinucleotides.
- CpG Islands and Promoter Regulation: The distribution of CpGs is not random. They are often clustered in regions known as CpG islands, which are frequently located near the transcription start sites (promoters) of genes.1 In actively expressed genes, these islands remain unmethylated. When these islands become heavily methylated, the gene is typically silenced.
- Mechanism of Repression: DNA methylation enforces silencing through two pathways. First, the bulky methyl group can physically impede the binding of transcriptional activators and RNA polymerase to the DNA. Second, and more importantly, methylated DNA recruits a family of proteins known as Methyl-CpG-binding domain (MBD) proteins (e.g., MeCP2). These proteins act as scaffolds to recruit additional repressive complexes, including histone deacetylases (HDACs) and corepressors, which condense the chromatin.4
- Genomic Stability: Beyond regulating gene expression, DNA methylation plays a critical defensive role. It serves to repress parasitic sequences, endogenous retroviruses (ERVs), and transposable elements such as LINE-1 and ALU repeats.5 By keeping these elements permanently methylated, the cell prevents their transcription and transposition, which could otherwise shatter the integrity of the genome.
1.2.2 Histone Modification: The Chromatin Scaffold
DNA in the nucleus does not float freely; it is wrapped around octamers of histone proteins to form nucleosomes, the fundamental repeating units of chromatin.5 The "tails" of these histone proteins protrude from the nucleosome core and are subject to a vast array of post-translational modifications (PTMs) that dictate the structural conformation of the chromatin.1
- The Histone Code: Specific combinations of modifications act as a code that "readers" (effector proteins) interpret to determine the state of a genomic locus.
- Acetylation: The addition of acetyl groups to lysine residues (e.g., H3K27ac) generally neutralizes the positive charge of the histone tails, reducing their attraction to the negatively charged DNA. This loosens the chromatin structure, creating a relaxed state known as euchromatin, which is accessible to transcriptional machinery.3
- Methylation: The effect of histone methylation depends on the specific residue and the degree of methylation. For example, trimethylation of histone H3 at lysine 4 (H3K4me3) is a hallmark of active promoters. In contrast, trimethylation at lysine 9 (H3K9me3) or lysine 27 (H3K27me3) recruits heterochromatin protein 1 (HP1) and Polycomb repressive complexes (PRC2), respectively, leading to the formation of tightly packed heterochromatin that is transcriptionally silent.3
- Interplay with DNA Methylation: There is frequently a reciprocal relationship between histone modifications and DNA methylation. For instance, H3K9me3 can recruit DNA methyltransferases to deposit methyl groups on the underlying DNA, reinforcing the silenced state.3
1.2.3 Non-coding RNA: The Guides
The third layer of regulation involves non-coding RNAs (ncRNAs), including microRNAs (miRNAs), long non-coding RNAs (lncRNAs), and small interfering RNAs (siRNAs).1 These RNA molecules do not code for proteins but play a pivotal role in guiding chromatin-modifying complexes to specific genomic loci.
- Mechanisms: ncRNAs can bind to DNA or mRNA transcripts to modulate expression. In the context of epigenetics, lncRNAs often act as scaffolds that bridge DNA and chromatin modifiers. A classic example is X-chromosome inactivation in female mammals, where the lncRNA Xist coats one of the X chromosomes and recruits repressive complexes to silence it entirely.7
- Therapeutic Potential: The ability of ncRNAs to target specific sequences makes them potent tools for regulating gene networks, a concept that is increasingly being integrated into therapeutic strategies alongside enzymatic editors.8
1.3 The Machinery of Maintenance: How Cells Remember
One of the most critical aspects of epigenetics for therapeutic application is the mechanism of maintenance. If epigenetic marks were lost every time a cell divided, any therapy relying on them would be transient and ineffective for chronic diseases. However, cells possess robust machinery to propagate these marks to daughter cells, a process termed epigenetic inheritance.
- DNMT1 and Maintenance Methylation: The primary enzyme responsible for maintaining DNA methylation patterns is DNA Methyltransferase 1 (DNMT1). During DNA replication, the double helix splits, and a new daughter strand is synthesized. This results in hemi-methylated DNA, where the parental strand carries the methyl group, but the newly synthesized strand does not. DNMT1 specifically recognizes these hemi-methylated sites and catalyzes the transfer of a methyl group to the corresponding cytosine on the new strand, thereby restoring the full methylation pattern.10 This fidelity ensures that a silenced gene remains silenced in the progeny.
- De Novo Methylation: In contrast, de novo DNA methyltransferases, such as DNMT3A and DNMT3B, are responsible for establishing new methylation patterns on unmethylated DNA.10 They are typically active during development or in response to specific signals. DNMT3L, a catalytically inactive cofactor, assists these enzymes by increasing their binding affinity and activity.10
- Active Demethylation: The removal of methylation is equally regulated, primarily by the Ten-Eleven Translocation (TET) family of enzymes. TET enzymes catalyze the oxidation of 5-methylcytosine to 5-hydroxymethylcytosine (5hmC) and further derivatives, which are eventually replaced by unmethylated cytosine through base excision repair pathways.13
Understanding this machinery is the prerequisite for understanding epigenetic editing: the technology essentially hijacks these natural "writers" (DNMT3A/3B), "readers" (MBDs), and "erasers" (TETs) to rewrite the cellular software.
Chapter 2: The Mechanics of Epigenetic Editing
2.1 From Observation to Manipulation
For years, epigenetics was an observational science; researchers correlated methylation patterns with disease states. The field transitioned to an interventional science with the development of programmable DNA-binding domains. The evolution of these tools parallels the history of gene editing itself.
Early attempts utilized Zinc Finger Proteins (ZFPs) and Transcription Activator-Like Effectors (TALENs) fused to epigenetic enzymes.14 While effective, these platforms were cumbersome to engineer, requiring the design of a new protein for every new DNA target. The revolution arrived with the adaptation of the CRISPR-Cas system.
2.2 The dCas9 Platform: A Dead Nuclease with a New Life
The workhorse of modern epigenetic editing is the "dead" Cas9 (dCas9) protein. In standard gene editing, Cas9 is a molecular scalpel that uses a guide RNA (gRNA) to find a specific DNA sequence and then cuts it with its two nuclease domains (RuvC and HNH).
To create an epigenetic editor, scientists introduce point mutations (typically D10A and H840A) into the nuclease domains of Cas9. This renders the protein catalytically inactive—it can still bind to DNA with high precision guided by the RNA, but it cannot cut.15 This transforms Cas9 from a pair of scissors into a programmable delivery vehicle.
By fusing effector domains to dCas9, researchers can deliver epigenetic payloads to any location in the genome.
2.3 The Toolkit of Effectors: Writers and Erasers
The functionality of an epigenetic editor is determined by the effector domain attached to the dCas9 anchor. These domains are drawn from the cell's natural regulatory toolkit.
2.3.1 Repressors (Silencing)
To turn off a gene, the dCas9 is typically fused to domains that promote heterochromatin formation and DNA methylation.
- KRAB Domain: The Krüppel-associated box (KRAB) domain is one of the strongest transcriptional repressors found in the human genome. When tethered to DNA, it recruits a complex known as KAP1, which in turn recruits histone deacetylases (like NuRD) and histone methyltransferases (like SETDB1) to deacetylate histones and deposit repressive H3K9me3 marks.16
- DNMT3A/3L: For more permanent silencing, the catalytic domains of DNMT3A and its cofactor DNMT3L are recruited. These enzymes deposit methyl groups directly onto the underlying CpG islands. The combination of KRAB (for initial chromatin compaction) and DNMT3A (for stable methylation) is a potent strategy for durable silencing, often referred to as CRISPRoff.4
2.3.2 Activators (Expression)
To turn on a gene, dCas9 is fused to domains that open chromatin and recruit transcriptional machinery.
- VP64/VPR: VP64 is a synthetic tetramer of the Herpes Simplex Virus VP16 activation domain. It recruits general transcription factors and chromatin remodelers. More potent versions, such as VPR (VP64-p65-Rta), combine multiple activation domains to drive robust expression.16
- p300/CBP: These are histone acetyltransferases (HATs) that deposit acetyl groups on H3K27, directly relaxing chromatin structure (euchromatin) and facilitating enhancer activity.16
2.3.3 Demethylators
To reactivate genes silenced by aberrant hypermethylation (e.g., tumor suppressor genes in cancer or FMR1 in Fragile X syndrome), dCas9 is fused to the catalytic domain of TET1. This enzyme converts methylcytosine to hydroxymethylcytosine, initiating the demethylation cascade and restoring gene expression.16
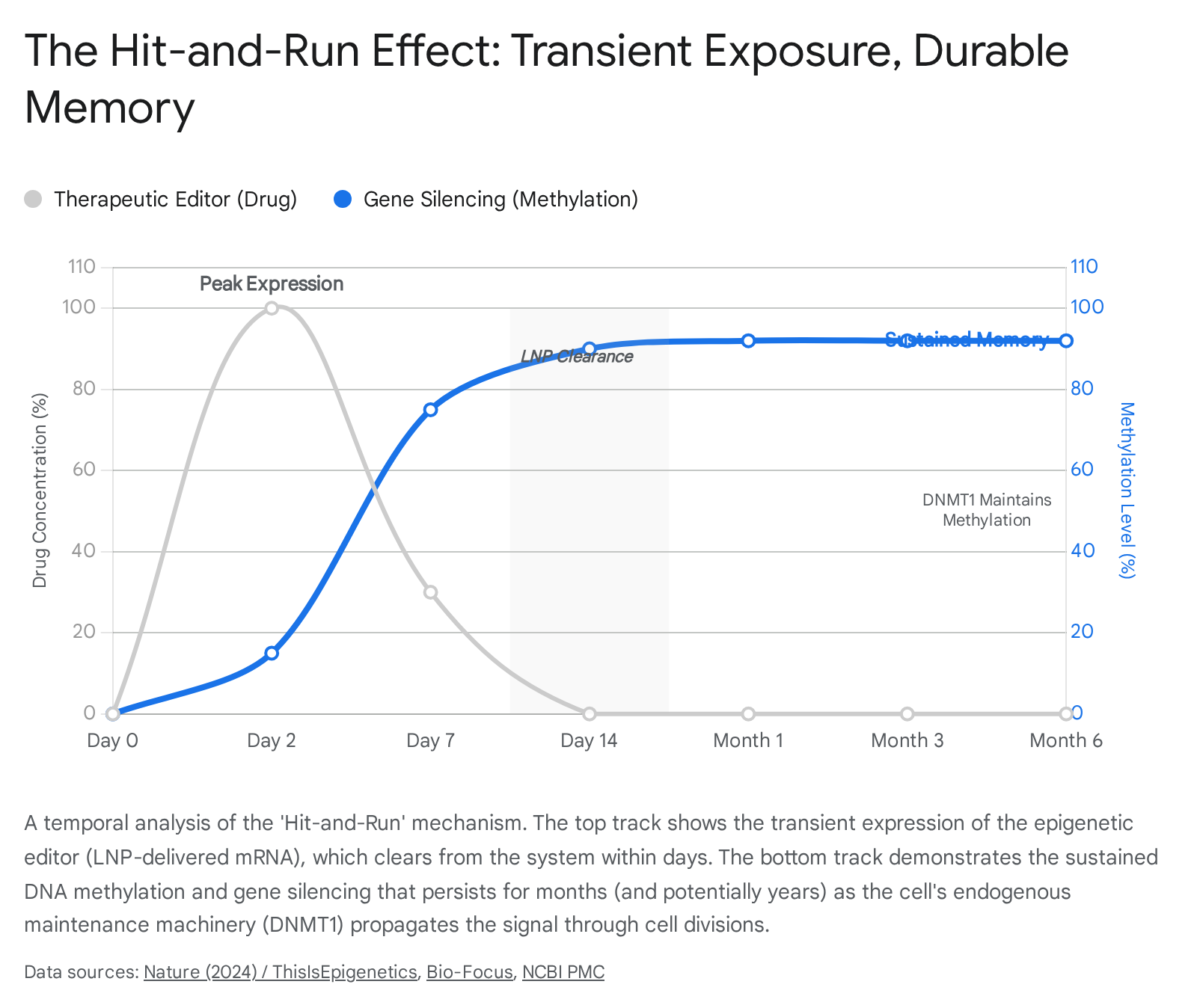
2.4 The "Hit-and-Run" Paradigm: Engineering Cellular Memory
A critical differentiator of epigenetic editing, and perhaps its most significant clinical advantage, is the potential for "hit-and-run" therapy. In traditional gene therapy, the therapeutic vector often needs to be permanently expressed to maintain the effect. In contrast, epigenetic editors can induce a state change that the cell then memorizes and propagates.
Research has demonstrated that transient delivery of epigenetic editors (e.g., via mRNA encapsulated in lipid nanoparticles) can establish a repressive heterochromatin environment that is self-sustaining. Once the de novo methyltransferases (DNMT3A/3L) establish high levels of CpG methylation and the local chromatin is compacted by KRAB-mediated mechanisms, the cell's endogenous maintenance machinery (DNMT1) recognizes these marks.10
When the cell divides, DNMT1 copies the methylation pattern to the daughter strands. Consequently, even after the original dCas9 editor has degraded and cleared from the system (the "run"), the gene remains silenced (the "hit"). This phenomenon was robustly demonstrated in landmark studies targeting the PCSK9 gene in mice. A single administration of an epigenetic editor resulted in stable gene repression and lowered cholesterol levels for nearly a year. Remarkably, this repression persisted even after partial hepatectomy forced the liver cells to regenerate and divide rapidly, proving that the epigenetic mark had been heritably encoded into the lineage.18
This durability without the permanent integration of foreign DNA or the permanent alteration of the host sequence is considered the "holy grail" of safety in genetic medicine, offering a potential functional cure with a finite treatment window.
Chapter 3: Comparative Analysis: Gene Editing vs. Epigenetic Editing
To understand the specific utility of epigenetic editing, it is essential to contrast it with the established modality of gene editing (CRISPR-Cas9). While both technologies share a targeting mechanism (RNA-guided Cas proteins), their approaches to modulating biology are fundamentally distinct, leading to divergent risk profiles and optimal use cases.
3.1 Mechanism: Hardware vs. Software Modification
- Gene Editing (CRISPR-Cas9): This approach functions by inducing Double-Strand Breaks (DSBs) in the DNA double helix. The cell's repair machinery—either Non-Homologous End Joining (NHEJ) or Homology-Directed Repair (HDR)—then rejoins the ends. NHEJ is error-prone and typically results in small insertions or deletions (indels) that disrupt the gene sequence, effectively "breaking" the gene to turn it off. HDR can be used to insert a corrected sequence, but it is inefficient in non-dividing cells.14 This is analogous to a "hardware" modification; the physical structure of the hard drive is altered.
- Epigenetic Editing: As described, this approach modulates the accessibility of the DNA without cutting it. It recruits the cell’s natural regulatory machinery to condense or relax chromatin. This is analogous to a "software" update; the hardware remains intact, but the permissions for reading the file are changed.2
3.2 Reversibility and Permanence
The permanence of gene editing is both its greatest strength and its most significant liability.
- Gene Editing: The changes made by CRISPR nucleases are permanent. Once a gene is knocked out or corrected, that alteration is inherited by all progeny cells forever. While this is desirable for curing a genetic defect, it also means that any off-target edit or unintended consequence is also permanent and potentially deleterious.21 There is no "undo" button for a double-strand break.
- Epigenetic Editing: Epigenetic modifications are theoretically reversible. While therapeutic designs aim for durability via DNA methylation, the marks can be removed by targeted administration of demethylases (e.g., TET enzymes) or may naturally fade over time if not sufficiently reinforced. This offers a potential safety valve: if an adverse event occurs, the therapy could theoretically be reversed or allowed to wash out, a feature not present in nuclease-based editing.1
3.3 Genotoxicity vs. Epigenotoxicity
The safety profile of the two technologies differs primarily in the type of damage they might cause.
- Genotoxicity (Gene Editing): CRISPR nucleases carry the risk of genotoxicity. This includes off-target cleavage (cutting similar sequences elsewhere in the genome), chromosomal translocations (where cut chromosomes fuse incorrectly), and large deletions. Furthermore, the creation of DSBs triggers the p53 DNA damage response, which can lead to cell cycle arrest or apoptosis and may select for p53-mutant cells, theoretically increasing cancer risk.14
- Epigenotoxicity (Epigenetic Editing): Epigenetic editors do not cut DNA, thereby eliminating the risks of translocations and DNA damage response. However, they introduce the risk of "epigenotoxicity." This involves off-target methylation, where the editor might transiently bind to a non-target CpG island and deposit methyl groups. While a single off-target cut can be catastrophic (e.g., disrupting a tumor suppressor), a small amount of off-target methylation is often tolerated by the cell unless it reaches a critical threshold that triggers heterochromatin spreading and gene silencing.24
3.4 The Multiplexing Advantage
One of the most profound advantages of epigenetic editing is in multiplexing—the ability to target multiple genes simultaneously.
- Gene Editing Limits: Attempting to cut multiple genes simultaneously with CRISPR nucleases is highly toxic. Multiple double-strand breaks can lead to "chromosomal shattering" (chromothripsis) or translocations, where the cut ends of different chromosomes join together. This toxicity severely limits the ability of gene editing to treat polygenic diseases that require the modulation of complex gene networks.26
- Epigenetic Solution: Because epigenetic editors do not cut the DNA, they do not threaten genomic integrity in the same way. It is theoretically possible to target dozens of loci simultaneously. A single therapeutic cocktail could "tune down" five different cholesterol-raising genes and "tune up" three protective metabolic genes. This capability to modulate gene networks rather than single switches is the key to treating complex, chronic diseases like cardiovascular disease, diabetes, and potentially psychiatric disorders.16
3.5 Summary of Differences
Feature | Gene Editing (CRISPR-Cas9) | Epigenetic Editing |
Primary Mechanism | DNA Double-Strand Breaks (DSBs) | Chromatin/Methylation Modulation |
Sequence Alteration | Yes (Permanent) | No (Preserves Sequence) |
Reversibility | Irreversible | Potentially Reversible |
Primary Risk | Genotoxicity (Indels, Translocations) | Epigenotoxicity (Off-target silencing) |
Heritability | Mendelian Inheritance | Variable (Subject to Reprogramming) |
Multiplexing | Difficult (Risk of toxicity/translocations) | Highly Feasible (Low toxicity) |
Delivery Payload | Nuclease + Guide | dCas9 fusion + Guide |

Chapter 4: The Safety Paradigm and Risk Assessment
While the "software" approach of epigenetic editing avoids the "hardware" damage of nucleases, it is not without its own unique set of risks. The safety discussion in the field is currently dominated by two primary concerns: the precision of the editing (off-target effects) and the potential for unintended inheritance (transgenerational effects).
4.1 Off-Target Epigenetic Remodeling and Detection
Just as Cas9 nucleases can cut the wrong DNA sequence if the guide RNA has partial complementarity to a non-target site, dCas9-methyltransferases can bind to near-match sequences and deposit methyl groups where they are not wanted.
- Mechanism of Error: The binding of dCas9 is governed by RNA-DNA base pairing. If the editor binds transiently to an off-target site, the fused methyltransferase (e.g., DNMT3A) may methylate nearby CpG dinucleotides. While a few misplaced methyl groups might not affect gene expression, the concern is "spreading." Epigenetic marks are cooperative; a small seed of methylation can attract binding proteins that recruit more methyltransferases, potentially causing heterochromatin to spread into a nearby promoter and silence a tumor suppressor gene or an essential metabolic gene.24
- Quantification and Surveillance: Detecting these errors is significantly more challenging than detecting DNA cuts. DNA cuts leave a scar (indel) that can be easily sequenced. Epigenetic changes are subtle. Detection requires advanced techniques like Whole-Genome Bisulfite Sequencing (WGBS) or ChIP-seq, which are data-intensive and expensive. However, recent comparative studies using Deep Learning models (like DeepCRISPR) and experimental data suggest that prime editing and epigenetic editors generally exhibit lower off-target rates than their nuclease counterparts, primarily because the "dwell time" required to establish a stable epigenetic mark is longer than the instant required to cut DNA, making transient off-target binding less functionally impactful.15
4.2 Transgenerational Epigenetic Inheritance (TEI): The Great Debate
One of the most contentious and theoretically significant risks is Transgenerational Epigenetic Inheritance (TEI). This refers to the possibility that epigenetic changes induced in an individual could be passed down to their offspring, affecting future generations who did not consent to the therapy.
- The Biological Barriers: In mammals, there are two massive waves of "global reprogramming" that act as firewalls against TEI.
- Zygotic Reprogramming: Shortly after fertilization, the paternal and maternal genomes are stripped of most methylation marks to creating a "clean slate" for the developing embryo (totipotency).30
- Primordial Germ Cell (PGC) Reprogramming: During the development of the sperm and egg cells in the embryo, a second wave of erasure occurs to reset imprinting marks.30
These barriers suggest that acquired epigenetic marks—including therapeutic ones—should be erased and not passed on.
- Evidence of Leakage: Despite these barriers, evidence exists that some marks can escape erasure. Historical studies on the "Dutch Hunger Winter" showed that nutritional stress in mothers affected the metabolism of their children and grandchildren, implying some form of inheritance.31 Animal studies have been more explicit; exposure of rats to endocrine disruptors (vinclozolin) induced methylation changes and fertility issues that persisted for four generations.31 Additionally, the Agouti mouse model has demonstrated that coat color changes linked to methylation status can be heritable.31
- Clinical Relevance: For the vast majority of current applications, this risk is minimal. Therapies targeting somatic tissues like the liver (via LNPs) or muscle (via AAVs) do not interact with the germline (sperm/eggs). The vector does not transduce germ cells. The risk is primarily theoretical for systemic delivery vectors that might cross the blood-testis barrier. Nonetheless, regulatory agencies like the FDA require rigorous assessment of germline transmission risks for all gene therapies.20
4.3 Immunogenicity
Another safety consideration is the immune response to the editor itself. The bacterial proteins used in these editors (Cas9 from Streptococcus pyogenes or Staphylococcus aureus) are foreign to the human body.
- Pre-existing Immunity: Many humans have pre-existing antibodies and T-cells against Cas9 due to common strep infections.
- Vector Immunity: The delivery vehicles, particularly AAVs, can also trigger neutralizing antibodies that render the therapy ineffective upon a second dose or cause inflammation.32
- Epigenetic Solution: While this is a shared risk with gene editing, epigenetic editing has a potential advantage. Because of the "hit-and-run" mechanism, the editor only needs to be expressed for a short window (days) to establish the memory. This contrasts with gene replacement therapies that require sustained expression. The transient nature of the treatment (e.g., using mRNA/LNP) reduces the duration of antigen exposure, potentially lowering the risk of a chronic immune response.34
Chapter 5: The Clinical Frontier and Industry Landscape
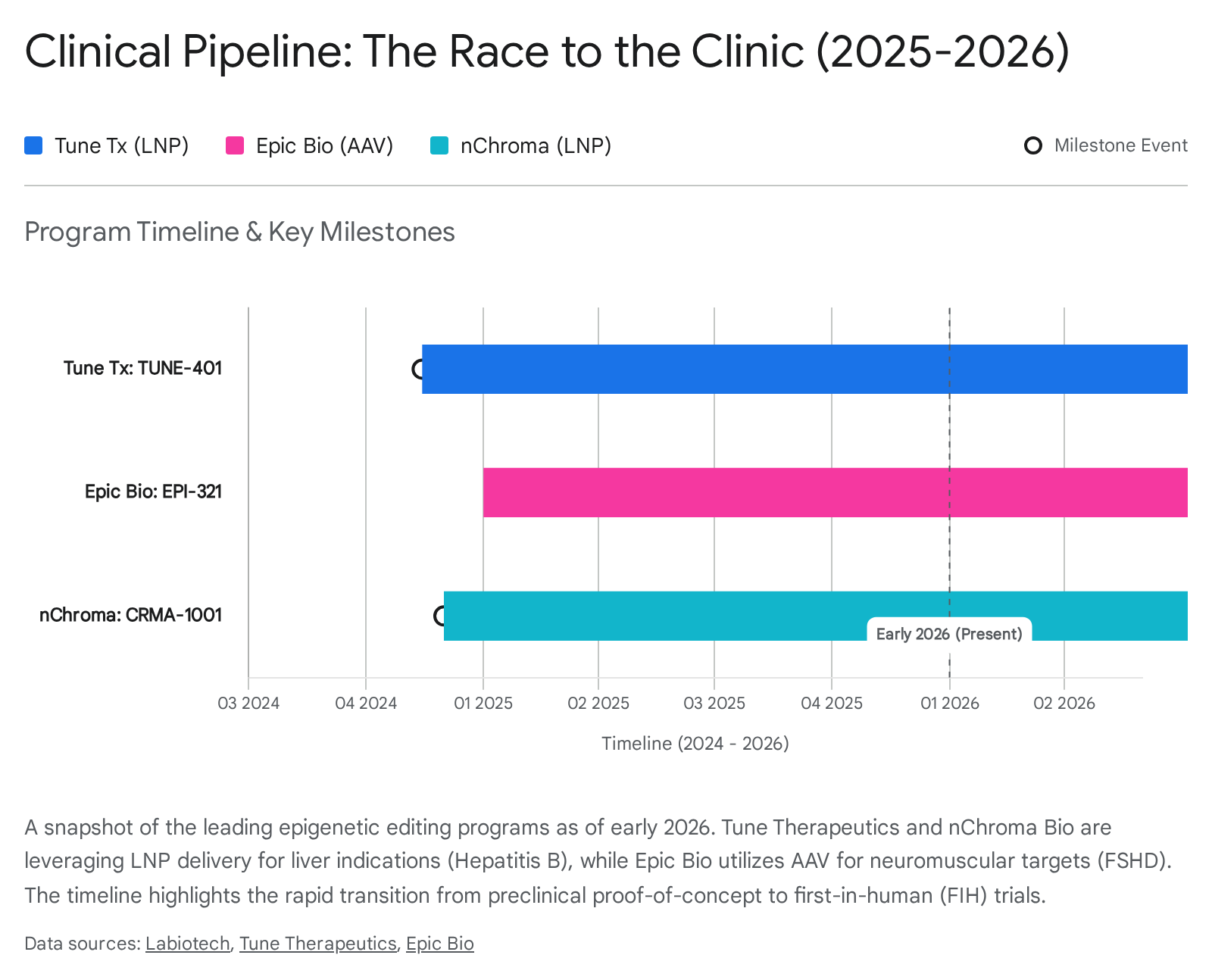
The transition of epigenetic editing from an academic curiosity to a clinical reality has been rapid. A cohort of specialized biotechnology companies has emerged, backed by significant venture capital, to translate these tools into medicines. The current clinical focus is strategically chosen to play to the strengths of the technology: targeting diseases where "tuning" is better than "breaking" and where delivery is feasible.
5.1 Tune Therapeutics: Viral Inactivation via Epigenetics
Tune Therapeutics, a leader in the space, has focused its proprietary "TEMPO" platform on a challenge that has baffled virologists for decades: Chronic Hepatitis B (HBV). Their lead candidate, TUNE-401, represents a novel application of epigenetic editing—targeting an exogenous viral genome rather than the host genome.
- The Clinical Unmet Need: Current standard-of-care treatments (nucleoside analogs) suppress active viral replication but fail to clear the virus. This is because the virus persists in the liver nucleus in two stable forms: covalently closed circular DNA (cccDNA) and integrated viral DNA. These act as a template for viral resurgence as soon as therapy is stopped.
- The TUNE-401 Mechanism: TUNE-401 utilizes lipid nanoparticles (LNPs) to deliver an epigenetic silencer that targets highly conserved regions of the HBV genome. It recruits methyltransferases to hypermethylate both the cccDNA and the integrated viral DNA.35
- The Data: In preclinical studies presented at AASLD, TUNE-401 demonstrated near-complete repression of viral DNA that was durable for over 550 days and through more than 275 cell doublings.35
- Trial Status: As of late 2024/early 2025, Tune Therapeutics received approval to initiate Phase 1b clinical trials in New Zealand and Hong Kong, marking the first time an epigenetic editor entered the clinic for an infectious disease.36
5.2 Epic Bio: Addressing Neuromuscular Disease
Epic Bio, leveraging its "GEMS" (Gene Expression Modulation System) platform, has taken a different approach, focusing on Facioscapulohumeral Muscular Dystrophy (FSHD).
- The Pathophysiology: FSHD is caused by the aberrant derepression of the DUX4 gene. In healthy adults, DUX4 is silenced by heavy methylation at the D4Z4 repeat region on chromosome 4. In FSHD patients, a contraction of these repeats leads to hypomethylation, allowing DUX4 to turn on. The DUX4 protein is toxic to muscle cells, causing progressive wasting.
- The Therapy (EPI-321): EPI-321 is designed to be a molecular "off switch." It uses a compact Cas-MINI protein (engineered to be smaller than standard Cas9) fused to epigenetic repressors. Its goal is to restore the missing methylation to the D4Z4 region, re-silencing the DUX4 gene and halting muscle toxicity.39
- Delivery Strategy: Unlike Tune's liver-targeted LNP approach, Epic Bio must reach muscle tissue. They utilize Adeno-Associated Virus (AAV) vectors, specifically serotype AAVrh74, which has high tropism for muscle.
- Clinical Status: Epic Bio initiated a first-in-human clinical trial for EPI-321 in 2025. The program has received FDA Fast Track and Orphan Drug designations, highlighting the regulatory support for this new modality.39
5.3 nChroma Bio: The Power of Consolidation
In a significant industry consolidation, December 2024 saw the merger of Chroma Medicine and Nvelop Therapeutics to form nChroma Bio. This merger illustrates the critical importance of delivery technology in the success of genetic medicines.
- The Rationale: Chroma Medicine possessed a leading epigenetic editing platform but faced the universal challenge of delivering these payloads to tissues outside the liver. Nvelop Therapeutics specialized in advanced delivery vehicles, including virus-like particles (VLPs) and non-viral systems. By merging, the new entity aims to package Chroma's editors into Nvelop's vehicles.42
- Lead Program: The merged company is prioritizing CRMA-1001, a liver-targeted epigenetic therapy for Hepatitis B and Delta virus. The company has explicitly stated a strategy of focusing on this lead liver asset while deprioritizing earlier-stage non-liver research to conserve capital and reach clinical milestones in 2026.44
5.4 The Delivery Bottleneck: LNP vs. AAV
The choice of delivery vehicle dictates the therapeutic focus of these companies. The field is currently split between two dominant modalities.
- Lipid Nanoparticles (LNPs):
- Pros: Non-viral, meaning low risk of genomic integration and lower immunogenicity. They can be manufactured at scale and allow for redosing (as they don't generate neutralizing antibodies against a capsid).
- Cons: When injected intravenously, they naturally accumulate in the liver due to ApoE-mediated uptake. While excellent for Hepatitis (Tune, nChroma) and cholesterol (PCSK9), targeting them to the brain or muscle is extremely difficult.32
- Adeno-Associated Virus (AAV):
- Pros: Excellent tissue tropism; specific serotypes can target muscle (AAVrh74), the eye, and the CNS.
- Cons: Small cargo capacity (~4.7kb) limits the size of the editor that can be packed. They are expensive to manufacture, and patients can have pre-existing immunity or develop antibodies that prevent a second dose. There is also a small but non-zero risk of viral integration into the genome.32
Chapter 6: Future Horizons and Emerging Applications
While the current clinical pipeline is focused on monogenic diseases and viruses, the unique capabilities of epigenetic editing—specifically multiplexing and partial reprogramming—position it to address challenges that are currently out of reach for traditional gene editing.
6.1 Multiplexing and Polygenic Diseases
The majority of the global disease burden—cardiovascular disease, Type 2 diabetes, Alzheimer's, psychiatric disorders—is not monogenic (caused by one gene) but polygenic (caused by the subtle interplay of hundreds of variants).
- The Limit of CRISPR: Traditional gene editing is ill-suited for these conditions. Cutting 50 genes simultaneously to "fix" a polygenic risk profile would be lethal to the cell, causing massive DNA damage and chromosomal chaos.26
- The Epigenetic Promise: Epigenetic editors do not break DNA. Therefore, it is theoretically possible to target dozens of loci simultaneously in a single cell without toxicity. A future therapy for heart disease might not just target PCSK9; it could simultaneously "tune down" five inflammatory genes and "tune up" three genes involved in lipid metabolism. This concept of "network medicine"—modulating entire pathways rather than single nodes—is the frontier of polygenic disease treatment.27 Recent advancements in "Multiplexed Orthogonal Base Editors" (MOBE) and similar epigenetic tools are paving the way for this reality.28
6.2 Anti-Aging and Partial Reprogramming
Perhaps the most ambitious and sci-fi application of epigenetic editing is the reversal of cellular aging. Aging is driven in part by "epigenetic drift"—the gradual loss of the precise methylation patterns that define a young, healthy cell.
- Reprogramming: In 2006, Shinya Yamanaka discovered that four transcription factors (OSKM) could reprogram any adult cell back into a stem cell, effectively resetting its age to zero. However, this process also erases cell identity (a neuron forgets it is a neuron), which leads to cancer (teratomas) if attempted in a living animal.45
- The Epigenetic Solution: Companies like Altos Labs (backed by billions in funding) and Retro Biosciences are pursuing "partial reprogramming." By using epigenetic controllers to transiently express reprogramming factors, they aim to reverse the epigenetic clock (rejuvenating the cell) without crossing the threshold of losing cell identity. This "epigenetic rejuvenation" could potentially restore the resilience of older tissues, treating age-related decline at its root cause.47
6.3 Immune Engineering
Epigenetic editing is also finding a home in oncology, specifically in the engineering of CAR-T cells.
- Problem: T-cells engineered to fight cancer often become "exhausted" over time, a state mediated by epigenetic silencing of effector genes and upregulation of checkpoint proteins like PD-1.
- Solution: Instead of knocking out PD-1 entirely (which can cause autoimmunity), epigenetic editors can be used to "dim" its expression or to program specific epigenetic states that resist exhaustion. This allows for the creation of "smart" immune cells with tunable potency and greater durability in the hostile tumor microenvironment.49
6.4 Agriculture and Biotic Stress
Beyond human health, epigenetic editing is making waves in agriculture. The ability to multiplex edits allows researchers to target complex traits like drought resistance or pathogen resistance in crops without introducing foreign DNA (transgenics) or causing genomic instability. Recent work has shown the feasibility of using CRISPR-based epigenetic tools to modify the promoters of susceptibility genes in plants, enhancing resistance to bacterial blight and other biotic stresses.26
Conclusion: The Software Update for Medicine
Epigenetic editing represents the maturation of the genomic revolution. If the first two decades of the 21st century were defined by the ability to "read" the genome (sequencing) and "cut" it (CRISPR), the coming decades will likely be defined by the ability to "control" it.
By decoupling gene regulation from DNA sequence modification, this technology solves the two greatest hurdles of traditional gene editing: safety (genotoxicity) and complexity (polygenic regulation). It offers a level of nuance—a dimmer switch rather than a breaker switch—that matches the biological reality of human disease.
The clinical data emerging from Tune Therapeutics and Epic Bio in 2025 and 2026 will be the critical proving ground. If the "hit-and-run" durability observed in animal models translates to humans, we may witness the emergence of a new class of "one-and-done" therapies for chronic viral infections, neuromuscular disorders, and potentially, the degenerative processes of aging itself. The future of medicine may not lie in rewriting our hard drive, but in optimizing the operating system that runs it.
Works cited
- Genetics, Epigenetic Mechanism - StatPearls - NCBI Bookshelf - NIH, accessed January 9, 2026, https://www.ncbi.nlm.nih.gov/books/NBK532999/
- Epigenetic Modification: Types, Functions, and Applications in Disease and Development, accessed January 9, 2026, https://www.cd-genomics.com/blog/epigenetic-modification-types-applications-diseases-development/
- Epigenetics - Wikipedia, accessed January 9, 2026, https://en.wikipedia.org/wiki/Epigenetics
- DNA methylation–independent long-term epigenetic silencing with dCRISPR/Cas9 fusion proteins | Life Science Alliance, accessed January 9, 2026, https://www.life-science-alliance.org/content/5/6/e202101321
- Epigenetics Modifications - DNA methylation | Proteintech Group, accessed January 9, 2026, https://www.ptglab.com/news/blog/epigenetics-modifications-dna-methylation/
- EZH2-DNMT1-mediated epigenetic silencing of miR-142-3p promotes metastasis through targeting ZEB2 in nasopharyngeal carcinoma - PMC - NIH, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC6748116/
- Epigenetic Modifications: Basic Mechanisms and Role in Cardiovascular Disease - NIH, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC3107542/
- Toward the Development of Epigenome Editing-Based Therapeutics: Potentials and Challenges - PMC - NIH, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC10003136/
- Epigenetic Regulation of Aging and its Rejuvenation - PMC - PubMed Central - NIH, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC12402629/
- Epigenetic editing: How cutting-edge targeted epigenetic modification might provide novel avenues for autoimmune disease therapy - PubMed Central, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC6078821/
- Epigenetic Interaction between UTX and DNMT1 Regulates Diet-Induced Myogenic Remodeling in Brown Fat | bioRxiv, accessed January 9, 2026, https://www.biorxiv.org/content/10.1101/2020.08.05.238923.full
- Inheritance of epigenetic transcriptional memory - PMC - NIH, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC10947848/
- Harnessing natural DNA modifying activities for editing of the genome and epigenome, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC6076857/
- Beyond the promise: evaluating and mitigating off-target effects in CRISPR gene editing for safer therapeutics - Frontiers, accessed January 9, 2026, https://www.frontiersin.org/journals/bioengineering-and-biotechnology/articles/10.3389/fbioe.2023.1339189/full
- Off-target effects in CRISPR/Cas9 gene editing - PMC - NIH, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC10034092/
- Recent Therapeutic Gene Editing Applications to Genetic Disorders - MDPI, accessed January 9, 2026, https://www.mdpi.com/1467-3045/46/5/255
- CRISPRoff epigenetic editing for programmable gene silencing in human cells without DNA breaks | bioRxiv, accessed January 9, 2026, https://www.biorxiv.org/content/10.1101/2024.09.09.612111v1.full-text
- From Code to Control: The Therapeutic Potential of Epigenome Editing | This is Epigenetics, accessed January 9, 2026, https://thisisepigenetics.ca/about-epigenetics/code-control-therapeutic-potential-epigenome-editing
- Hit-and-run Epigenome Editing: Gene Therapy Advancements - BioFocus, accessed January 9, 2026, https://www.bio-focus.co.uk/biotechnology/hit-and-run-epigenome-editing%3A-gene-therapy-advancements
- Comparative ethical evaluation of epigenome editing and genome editing in medicine: first steps and future directions - NIH, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC11137457/
- The Issue of Inheritance in Epigenome Editing - Technology Networks, accessed January 9, 2026, https://www.technologynetworks.com/genomics/articles/the-issue-of-inheritance-in-epigenome-editing-382121
- CRISPR Off-Target Effects: Mechanisms and Solutions | Danaher Life Sciences, accessed January 9, 2026, https://lifesciences.danaher.com/us/en/library/crispr-off-target-effects.html
- Mitigating off-target effects in CRISPR/Cas9-mediated in vivo gene editing - PubMed Central, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC7220873/
- Detecting Methylation Changes Induced by Prime Editing - PMC - NIH, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC12294443/
- Occurrence and Nature of Off-Target Modifications by CRISPR-Cas Genome Editing in Plants | ACS Agricultural Science & Technology, accessed January 9, 2026, https://pubs.acs.org/doi/10.1021/acsagscitech.1c00270
- Turbo-charging crop improvement: harnessing multiplex editing for polygenic trait engineering and beyond - UGA Plant Center, accessed January 9, 2026, https://plantcenter.uga.edu/turbo-charging-crop-improvement-harnessing-multiplex-editing-for-polygenic-trait-engineering-and-beyond/
- Modeling the complex genetic architectures of brain disease - PMC - NIH, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC7909729/
- Gene Editing Breakthrough: Multiplex Genome Editing for Complex Diseases - SynBioBeta, accessed January 9, 2026, https://www.synbiobeta.com/read/gene-editing-breakthrough-multiplex-genome-editing-for-complex-diseases
- Full article: CRISPR-Based Epigenome Editing: Mechanisms and Applications, accessed January 9, 2026, https://www.tandfonline.com/doi/full/10.2217/epi-2023-0281
- Transgenerational Epigenetic Inheritance: myths and mechanisms - PMC - PubMed Central, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC4020004/
- Full article: Calling the question: what is mammalian transgenerational epigenetic inheritance? - Taylor & Francis Online, accessed January 9, 2026, https://www.tandfonline.com/doi/full/10.1080/15592294.2024.2333586
- Epigenetic Editing in Neurological and Neuropsychiatric Disorders: Pioneering Next-Gen Therapeutics for Precision Gene Control - PubMed Central, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC12743672/
- Trends and challenges of AAV-delivered gene editing therapeutics for CNS disorders, accessed January 9, 2026, https://pmc.ncbi.nlm.nih.gov/articles/PMC12341529/
- The next frontier of in vivo gene therapy: are LNPs the new AAV? - Dark Horse Consulting, accessed January 9, 2026, https://www.darkhorseconsultinggroup.com/post/the-next-frontier-of-in-vivo-gene-therapies
- Tune Therapeutics Presents First Data Supporting TUNE-401: a First-in-Class Epigenetic Silencer for Hepatitis B, accessed January 9, 2026, https://tunetx.com/tune-therapeutics-presents-first-data-supporting-tune-401-a-first-in-class-epigenetic-silencer-for-hepatitis-b/
- Tune Therapeutics Moves into Clinical Spotlight with TUNE-401: A First-in-Class Epigenetic Silencer for Hepatitis B, accessed January 9, 2026, https://tunetx.com/tune-therapeutics-moves-into-clinical-spotlight-with-tune-401-a-first-in-class-epigenetic-silencer-for-hepatitis-b/
- AASLD 2024: Tune Therapeutics Shows Near-Complete HepB Repression With Tune-401 Epigenetic Silencer, accessed January 9, 2026, https://tunetx.com/aasld-2024-tune-therapeutics-shows-near-complete-hepb-repression-with-tune-401-epigenetic-silencer/
- Tune Therapeutics Receives Approval to Initiate Clinical Trial in Hong Kong for HepB Epigenetic Silencer, accessed January 9, 2026, https://tunetx.com/tune-therapeutics-receives-approval-to-initiate-clinical-trial-in-hong-kong-for-hepb-epigenetic-silencer/
- Epicrispr Biotechnologies Secures $68 Million Series B to Initiate Clinical Trial for First-in-Class Disease-Modifying Epigenetic Neuromuscular Therapy for FSHD, accessed January 9, 2026, https://epicrispr.com/epicrispr-biotechnologies-secures-68-million-series-b-to-initiate-clinical-trial-for-first-in-class-disease-modifying-epigenetic-neuromuscular-therapy-for-fshd/
- Epicrispr Biotechnologies Presents Data on Powerful Gene Reactivation with GEMS Epigenetic Editors at ESGCT, accessed January 9, 2026, https://epicrispr.com/epicrispr-biotechnologies-presents-data-on-powerful-gene-reactivation-with-gems-epigenetic-editors-at-esgct/
- Epicrispr Reports Early Clinical Activity and Favorable Safety Profile in First-in-Human Epigenetic Editing Study for FSHD, accessed January 9, 2026, https://www.fshdsociety.org/2026/01/08/epicrispr-reports-early-clinical-activity-and-favorable-safety-profile-in-first-in-human-epigenetic-editing-study-for-fshd/
- Chroma Medicine and Nvelop Therapeutics Unite to Form nChroma Bio, Securing $75 Million to Accelerate Genetic Medicines, accessed January 9, 2026, https://www.nchromabio.com/press-release/chroma-medicine-and-nvelop-therapeutics-unite-to-form-nchroma-bio-securing-75-million-to-accelerate-genetic-medicines/
- Chroma Medicine and Nvelop Tx merge to launch nChroma Bio with $75M - BioXconomy, accessed January 9, 2026, https://www.bioxconomy.com/investment/chroma-medicine-and-nvelop-tx-merge-to-launch-nchroma-bio-with-75m
- Genetic medicine outfit nChroma sheds staff in research reorg - Fierce Biotech, accessed January 9, 2026, https://www.fiercebiotech.com/biotech/genetic-medicine-outfit-nchroma-sheds-staff-research-reorganization
- 13 anti-aging biotech companies leading longevity in 2025 - Labiotech.eu, accessed January 9, 2026, https://www.labiotech.eu/best-biotech/anti-aging-biotech-companies/
- Science - Altos Labs, accessed January 9, 2026, https://www.altoslabs.com/science
- 9 Anti-Aging and Longevity Startups to Watch in 2026 - Cure., accessed January 9, 2026, https://wewillcure.com/insights/company-profiles/anti-aging-and-longevity-startups-to-watch
- The business of longevity in 2025: big bets amid biotech bust, accessed January 9, 2026, https://thelongevityinitiative.org/2026/01/business-2025-bets-biotech-bust/
- Epigenetic editing enables safer and more effective T cell therapies - News-Medical.Net, accessed January 9, 2026, https://www.news-medical.net/news/20251021/Epigenetic-editing-enables-safer-and-more-effective-T-cell-therapies.aspx
- Multiplex Genome-Editing Technologies for Revolutionizing Plant Biology and Crop Improvement - Frontiers, accessed January 9, 2026, https://www.frontiersin.org/journals/plant-science/articles/10.3389/fpls.2021.721203/full
